DNA Strand
General
- Submit all files to elearn. Make sure to name file(s) EXACTLY as specified - including capitalization. Do not zip or otherwise package/compress files unless told to.
- Make sure your program accepts input in EXACTLY the order and format given.
Do not ask for extra input or change the order of inputs.
If you want to make an expanded version of a program that and get feedback on it, email it to me or show me in class. - Slight variations in output wording/formatting are usually fine
(as long as formatting isn't part of the assignment).
If you do not get the right final output, print out some evidence of the work your program did to demonstrate your progress. - Readability and maintainability of your code counts. Poor formatting, confusing variable names, unnecessarily complex code, etc… will all result in deductions to your score.
Background
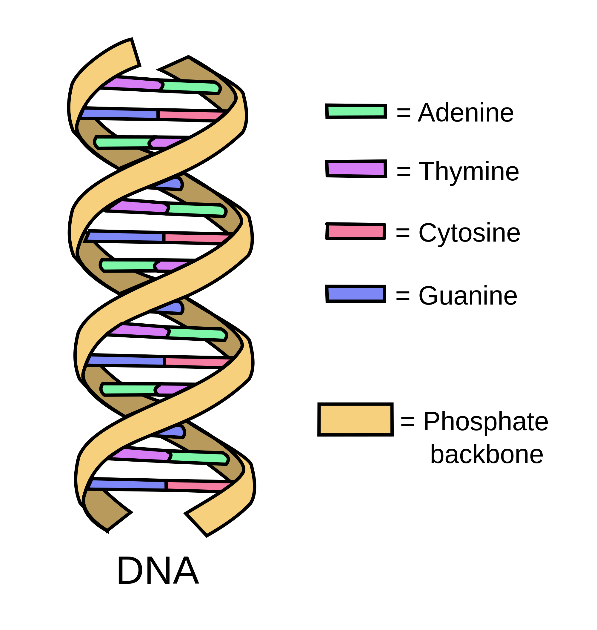
DNA, or deoxyribonucleic acid, is the primary carrier of genetic information in most organisms. The information in DNA is represented using a string of nucleotides. There are four kinds of nucleotides: Adenine, Thymine, Cytosine, and Guanine. They are typically abbreviated to their initials, so a strand of DNA might be expressed as "ACTTGAT".
These strands join together with a complementary strand to form
DNA's famous double helix shape. Adenine and Thymine always pair
up and Cytosine and Guanine always pair up. Thus, the strand:ACTTGAT
Would always have a complementary strand:TGAACTA
Where A in the original lines up with T in the new strand, C in the
original matches G in the new strand, etc…
Assignment Instructions
Submit file: assign5.zip
Make sure to compress your entire folder, not just your code file.
I should be able to add my own copies of DNAStrand.h and doctest.h and build your project with the following command:
g++ -g -std=c++17 DNAStrand.cpp tests.cpp -o tests.exe
Your task is to make a class that represents a DNA strand. A DNAStrand will track its length and an array of Bases. Base is defined as an enumerated type inside DNAStrand with the possible values A, C, G, and T).
We will provide for creating a DNAStrand from a string, like this:
DNAStrand strand1("ACTGAGATA");
But the data will always be stored as bases. The starter code has some
simple functions to convert between chars like 'T' and Bases like T.
DNAStrand will also provide various operators and functions for doing things like combining two strands, getting the complement of a strand or searching for one strand inside another.
You are provided a DNAStrand.h, a partial DNAStrand.cpp, and tests.cpp. You should not modify DNAStrand.h. You should only modify tests.cpp to comment out entire TEST_CASEs that do not compile with your code or that cause a crash. Do not modify a TEST_CASE in any way except to comment the entire thing out. Leave in any TEST_CASES that compile and run, even if they fail.
Passing Test > Failing Test > Commented-Out Test > Does not Compile
See below for tips on specific functions.
Getting Started
You should make a UnitTest project and add the provided files to it,
replacing the original tests.cpp.
Update the Makefile so that the HEADERS and TEST_FILES list
the .h and .cpp files respectively.
# list .h files here
HEADERS = DNAStrand.h
# list .cpp files here
TEST_FILES = tests.cpp DNAStrand.cpp
Comment out all the tests but the first one in tests.cpp. Implement the constructor and work until you pass test 1. Then, work on one new test at a time, implementing only the new functions needed to pass that test.
Memory Management
Make sure to test your code for leaks or other memory errors with Valgrind (or leaks on a Mac).
Test early and often. It will be much easier to find and fix memory issues write after you write the relevant code than if you wait until you have written all the functions and then test with Valgrind (or leaks on a Mac) for the first time. Valgrind should give your program a clean bill of health. Any errors reported by Valgrind will result in a significant deduction, even if the program appears to work fine despite of them.
In particular, watch out for memory leaks. Memory leaks will not cause any visible errors in your program—the only way to know that you have a leak is to use Valgrind (or run your code long enough that you can measure a steady increase in used memory).
Remember, Valgrind shows you where it detected an issue. Often, the place where you caused the issue is somewhere else! For example, maybe your constructor did not allocate an array but that was not a problem until later when you went to write to that array.
Function Tips
Here is a guide to function implementations. They are provided in a suggested implementation order.
Design Note:
Most of the functions we will write are non-modifying - they
produce a new DNAStrand instead of modifying the existing one.
However, just because you are making a new DNAStrand does not mean you
need to use the new keyword. Allocate the "new" objects on the stack as
local variables and return them. The new keyword should only be used
for allocating the memory used by a DNAStrand (in its constructor).
baseToChar and charToBase are provided as static members
of DNAStrand. Use them any time you need to change an enumerated Base value
into a char or vice verse.DNAStrand(const string& startingString)Our basic constructor — it should create a DNAStrand that represents the same sequence of bases as
startingStringcontains.A DNAStrand's
basesshould be used to track a dynamically allocated array that is the exact size needed to store the strand.There is no magic way to convert the entire string into the array of bases. You will need to set the bases in your array one by one to correspond to the characters in the string.
~DNAStrand()Destructor. Not tested by the unit tests, but every test will leak memory without this. Verify it works by running your program with Valgrind (or leaks on a Mac).
DNAStrand::Base at(int index) constThis should return the specified particular base from the strand.
Because the return type is not inside the function, we have to specify
DNAStrand::Base. Inside the function itself, or any other member functions, you can just refer to the Base type as aBase.This needs to throw an
out_of_rangeexception if given a bad index (less than 0 or past the end of the array).string toString() constThis should turn a strand like
ACCTinto the text"ACCT". There is no magic shortcut to turn you array of bases into a string. You will have to build the result string up one character at a time.Overriding the
<<operator is the more idiomatic C++ way of doing output:cout << dna;instead ofcout << dna.toString();If you want, feel free to experiment with it in a copy of the project. But since you can't modify the .h file, you can't substitute it for toString in the version you turn in.bool operator==(const DNAStrand& other) constChecks to see if the strands are the same length and contain an identical list of bases. (i.e.
"ATTAG"and"ATTAG"). Needed by the first test (and most others).Warning: you can't just compare the
basesarrays themselves with==; that will compile, but it will just compare the memory addresses of the two arrays. You need to loop through thebasesto compare the individual Bases.DNAStrand(const DNAStrand& other)andDNAStrand& operator=(const DNAStrand& other)Copy constructor and assignment operator. Tested in two unit tests. These are another big area for memory issues — test with Valgrind (or leaks on a Mac)!
The tests for many other functions will rely on the copy constructor. You need to get it working before finishing most of the rest of the assignment.
DNAStrand(int lengthValue)This private constructor builds a DNAStrand with the given length and should allocate an array big enough to hold that many Bases Having this constructor will make other functions easier to write as they will be able to do something like:
DNAStrand temp(20); // I know it should be 20 long //use loop to set temp's bases return temp;It is private as we don't want other code using this special helper constructor. There are no tests for this (since it is private). But you will probably want to use it as part of any function that returns a DNAStrand.
operator+andgetComplementCheck the .h for documentation. Note that both return a new object that has the desired DNAStrand. Don't try to modify the current strand. You will probably need/want the DNAString(int length) constructor as a helper for these.
See the note above about "new" vs
new! You probably should not be usingnewin this function.Each has their own test.
DNAStrand substr(int start, int substrLength) constGet a new DNAStrand that contains the indicated portion of this DNAStrand. This should throw an
out_of_rangeexception if the index or substrLength are negative, or if the start + substrLength would go past the end of the strand.You will probably need/want the DNAString(int length) constructor as a helper for this!
As a student of computing science, you should notice that we are being a bit wasteful with storage in this design. With four possibilities for each nucleotide, 2 bits are sufficient to distinguish which one we mean. However, an enum is typically stored with 32 bits meaning that each Base in this program occupies 4 bytes.
If storage were a concern, we could fit 4 nucleotides per byte, at the expense of the code to read and write them being more complicated. This would reduce the storage for our data by a factor of 16.